Artificial intelligence gets a lot of scare stories in the media at the moment. But AI is a powerful tool that can be used for good as well, especially in fields that now generate tons of data, like medicine.
Artificial intelligence gets a lot of scare stories in the media at the moment. But AI is a powerful tool that can be used for good as well, especially in fields that now generate tons of data, like medicine.
One of the big challenges in modern medicine is keeping ahead of the microbes that cause infectious diseases. Can AI, in particular a small part of AI called machine learning, help us detect and stop deadly infectious diseases before they spread and become harmful epidemics?
These bugs evolve quickly. Any mutations they develop that help them become more infectious will spread rapidly throughout a population.
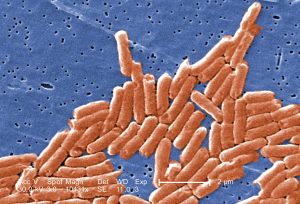
For example, the bacteria known as Salmonella. Most Salmonella live happily in our guts, and don’t cause too many problems. They might cause food poisoning occasionally, but they won’t kill you. Doctors and researchers who study Salmonella call this type ‘gastrointestinal’ because it doesn’t move out of the digestive tract. But sometimes these gastrointestinal bacteria mutate and evolve into really nasty bugs, that make us very sick and are much more likely to kill us, like the type of Salmonella that causes typhoid fever or meningitis. These are called invasives.
When a person turns up to a hospital and they are sick with a Salmonella infection that hasn’t been seen anywhere else, it is hard to know whether their infection is gastrointestinal or invasive.
Dr Paul Gardner, who recently joined the Otago Department of Biochemistry, and his colleagues Dr Nicole Wheeler, from the Wellcome Sanger Institute, and Dr Lars Barquist, from the Helmholtz Institute for RNA-based Infection Research in Germany, put their minds to this challenge.
They wanted to know if they could tell whether a Salmonella infection is the less harmful type or a really nasty one that cause life-threatening epidemics, simply by looking at its DNA sequence.
The team is particularly interested in using DNA mutations that appear in genes that code for proteins.
What exactly are proteins again?
Proteins carry out nearly all of the tasks that have to be done in a cell – building things up, breaking things down, moving things around and providing structural support.
Nearly everything in a cell is made from, or by, proteins. There are millions of different proteins, each with a different job and a different shape. One of the most important things that DNA does is carry the instructions for making proteins, in the form of genes.
If you want an introduction to proteins, this video from Learn.Genetics is a good start. https://learn.genetics.utah.edu/content/basics/proteins/
So the researchers look at mutations in protein genes?
Yes, Paul and his colleagues focus on finding mutations from DNA sequence data, in particular the ones that occur in the conserved parts of a protein.
When a species evolves, over a long time their proteins gradually change.
But not all proteins change at the same rate. And even different parts within the same protein will change at different rates.
The parts of a protein that are really important for doing its job change very slowly, sometimes not at all for a very long time, so we say that they are conserved. If a change/mutation does occur in this part, scientists get really interested because it means the protein might be changing how it works – possibly because the creature it belongs to has to get used to a change in its environment.
By looking at the mutations in a conserved bit of a protein, the researchers can predict whether the protein will have changed its function a little or a lot, and score it accordingly, without even knowing what the protein does.
But which mutations, in which genes, are the ones that you see in the nasty Salmonella vs the not so nasty?
Good question. You could try to look at each mutation in each individual gene, in the nasty invasive Salmonella, compare these to what you see in gastrointestinal Salmonella, and then come up with some general rules according to what you found. But there are potentially hundreds of mutations per gene, and thousands of genes to look at in each Salmonella strain. My head hurts just thinking about it.
Instead, Paul, Nicole and Lars got a clever computer algorithm on the job. Using a type of machine learning called a “Random Forest” approach, they made a tool that analyses which mutations make Salmonella turn nasty. They trained the tool using previously sequenced groups of Salmonella, including six Salmonella strains that caused invasive infections in the past, and seven gastrointestinal strains.
Technically they weren’t using an AI for this work. Machine learning is a small subset of AI – basically they taught a machine to do one job very well, whereas with AI it would be able to solve larger more complex problems. Machine learning is like AI’s little sister. Kind of like the difference between a neuron and a brain.
At first the tool looked at 6,438 genes across those 13 strains. But they were able to whittle that down. In the end, only 196 of the genes were needed to predict invasiveness in Salmonella. Perhaps unsurprisingly, a lot of these key genes turned out to be involved in metabolism.
So they made an ‘invasive Salmonella prediction tool’, but does it work?
The team tested the tool on genomic DNA sequence data collected from hundreds of different Salmonella strain samples collected from around the world.
The tool gave each strain an ‘invasiveness index’ score, based on whether the mutations found in those strains were like the mutations seen in the invasive strains or the mutations in the gastrointestinal strains during the Random Forest training.
The results were great. It correctly sorted out the invasive Salmonella strains from those associated with gastrointestinal infection.
So where to now?
Doctors and scientists are now starting to sequence the whole genomes of microbes on a routine basis as part of monitoring the spread of infectious diseases, generating massive DNA data sets as they go.
Paul is excited about how this tool can be used with that data. “Pretty soon that technology will be accessible to even country hospitals, and places like that will be routinely sequencing anyone who looks like they might be carrying some sort of infectious disease.”
“So we can use our methodology, attach it to these new sequencing devices and characterise what is actually causing the disease. Because at the moment that’s still a very difficult thing to establish. Hopefully in the very near future we’ll be able to have a very quick turnaround on what’s causing disease.”
He is also optimistic that the methods they have used here could also be used in the fight against growing bacterial resistance to antibiotics.
“We’ve looked at mutations in bacteria that switch from a relatively benign lifestyle (i.e. may cause food poisoning) to an invasive lifestyle (i.e. may cause typhoid fever and death). We found that we can predict, [using] just genome sequence, which lifestyle the bacteria [are] adapting to.”
“In the future we may be able to apply a similar approach to the antibiotic resistance problem.”
Read the full paper here:
Machine learning approach employed to identify signatures of host adaptation in the bacterial pathogen Salmonella enterica. Nicole E. Wheeler, Paul P. Gardner, Lars Barquist. PLoS Genetics. https://doi.org/10.1371/journal.pgen.1007333