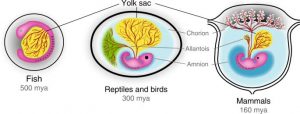
Figure 1 – Amniotic Eggs vs Placental Embryos. The advent of the placenta lead to the regression of the yolk and allantois, with the obvious loss of the calcified shell and retention of the amniotic sac. The mammalian placenta is estimated to have evolved between 130-160 million years ago, which is near the dawn of the mammalian phylogenetic class [7 & 9].
What is the placenta?
It is a supportive organ of embryogenesis that it is extensively developed and diverse in eutherian mammals. When compared to the yolk sac (‘nutrient reserve’) and allantois (‘waste tank’) of fish, amphibian, reptilian and avian eggs, the placenta allows for much greater exchange of nutrients and waste [2, 6, 8 & 9]. This enables the eutherian embryos to develop for longer and grow larger within the uterus. However, this benefit is traded off for the potential harms that can emerge from internal reproduction e.g. – spontaneous abortion (miscarriage) and/or severe haemorrhaging/infection during delivery [2, 6, 8 & 9]. In this way, the placenta evolved as a mechanism to increase the reproductive investment from a mother towards their offspring, allowing the offspring to have greater fitness when born. This supportive organ was derived from the trophoblasts that compose the extraembryonic membranes of a blastocyst [2, 6, 8 & 9]. These membranes initially evolved to encase the embryo in protective amniotic sac that has a yolk, allantois and shell. During evolution the trophoblasts of the embryonic membranes were transformed via endo-retroviral genes (ERGs) to seek out and implant themselves into the maternal decidua. Afterwards, they invaded the inner linings of the uterine wall to access the maternal circulation, so it can become a novel source of nutrients and dump for waste [2, 6, 8 & 9].
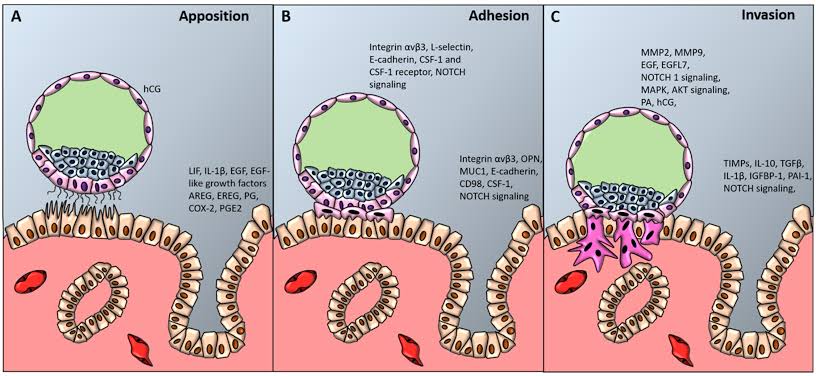
Figure 2 – Blastocyst Implantation into Maternal Decidua. The Inner Cell Mass (ICM) are the blue cells, and the Outer Cell Mass (OCM) are the pink cells, of the blastocyst. Apposition is where the blastocyst orientates itself to properly land on endometrial epithelium, with the ICM facing towards the maternal decidua [6]. This stage is facilitated by the microvilli of the trophoblasts in the OCM, which get attached to the pinopodes of the endometrial epithelium. Adhesion is where integrins/selectins on the trophoblasts and maternal epithelium interlock to form a stronger attachment [6]. Invasion is where the trophoblasts differentiate into syncytotrophoblasts and cytotrophoblasts. The syncytotrophoblasts form the multinucleated finger-like projections of the blastocysts to burrow through the maternal decidua, and they secrete proteolytic and hydrolytic enzymes to break down the maternal blood vessels [6]. In addition, the syncytotrophoblasts secrete suppressive chemokines like IL-10 to prevent maternal rejection of the invading blastocyst. The cytotrophoblasts act like stem cells that consistently generate new syncytotrophoblasts, and they direct the embryoblasts of the ICM to build the chorionic blood vessels from mesenchymal cells [6].
A blastocyst is mass of 200-300 cells that were generated from a fertilized egg, and it contains the ICM and the OCM compartments. The ICM creates the germ cell layers which constitute the embryoblasts; whereas the OCM form the trophoblast cell layers that make up the placenta [6, 8 & 9]. Placentas cultivate a signaling network of hormones and chemokines with the maternal body to ensure that there is safe gestation (pregnancy). Many complications during pregnancy can be traced to a placenta that has dysregulated hormone signals, malformed blood vessels and/or chemokine over-reactivity [2, 6, 8 & 9]. In this case, dysregulated hormone signals can be excessive insulin from the placenta that causes pregnancy-related diabetes; whereas, malformed blood vessels in the intervillous space of the placenta can cause preeclampsia – which is restricted circulation for the growing foetus and high blood pressure for the mother. In addition, chemokine over-reactivity could manifest as the placenta producing excessive peptides that are paternally derived, which can have adverse reactions with the maternal immune system – leading to placental inflammation and/or foetal calcification [6, 8 & 9].
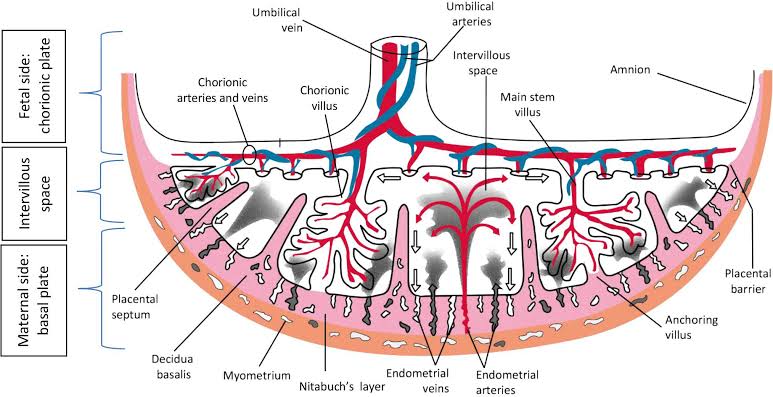
Figure 3 – Discoid Placenta Macro-anatomy. Placentation is driven by highly conserved transcriptional programs that are expressed in trophoblasts. This transcriptional program is transient and responsive to the maternal decidua [5 & 8]. It allows the finger-like syncytotrophoblasts projections to carve out lacunae in the maternal decidua, which mark the boundaries of the primary, secondary and tertiary villi. In this case, the lacunae are the circulating pools of maternal blood that are exchanging metabolites and cellular signals with the villi of the human placenta [5 & 8]. The cytotrophoblasts during gestation will direct the continued expansion and folding of the villi, where in the human placenta they generate between 16 to 20 lacunae units that have a 12 meter-squared surface area [5 & 8].
The sex-specific transcriptomes that arise during placental development emerge from various genetic elements, ranging from chromosomal complement (CC), to pseudo-autosomal regions (PARs), X-inactivation escapees (XEs), sex-biased epigenetic marks (SEMs) and hormone response elements (HREs). The chromosomal complement is the set of sex chromosomes that you receive from your parents, where XY is stereotypically male and XX is canonically female [1 & 3]. The male CC would give you a dose of X- and Y-linked genes, whereas, a female CC you would only get you a dose of X-linked genes. The PARs are sequences of DNA that are homologous between the X and Y chromosomes – where males would have X and Y PAR alleles, and females would only have X PAR alleles [1 & 3]. The XEs are regions of DNA that have not been silenced by the process of X-inactivation. The XEs would mean that females would have 2 doses of genes that are expected to have only one expressive allele [1 & 3]. The SEMs can be methylation (silencing) or acetylation (expressive) marks that are more prevalent in the chromatin landscape of a specific sex, where there would be distinct epigenetic profiles for males and females. The HREs are genes that are expressed or silenced in response to either estrogen or testosterone signaling pathways – this is more relevant in 2nd and 3rd trimester of human gestation [1 & 3].
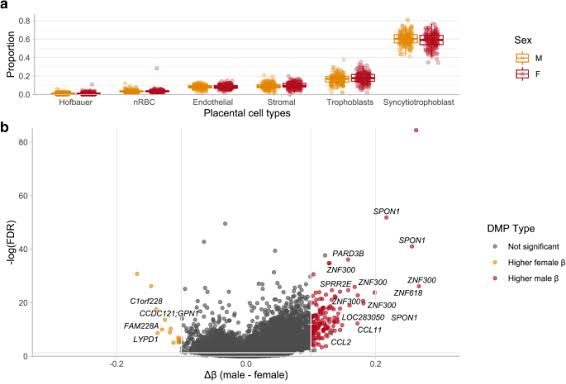
Figure 4 – Counts for Placental Cell Types in part A; Volcano Plot for Differentially Methylated CpG Sites in part B. Part A illustrates that the male and female placentas contain the same amount of all placental cell types. Part B shows that the female placenta has 146 CpG sites out of 162 that are methylated [4]. This would mean that in the male placenta woud have146 CpgG sites that are unmethylated, and they sit 100 bp away from gene promoters, so it would results in 146 autosomal genes being upregulated in male placenta. Most of these upregulated autosomal genes were revealed by gene ontology to be involved with chemotaxis and chemokine signaling – which suggests that male placentas would be more reactive to immunological stressors from the maternal circulation than female placenta [1, 3 & 4].
Male syncytotrophoblasts exhibit increased expression of chemokine TNF-α and NF-κB. These are thought to produce a faster adhesion stage during blastocyst implantation [1, 3 & 4]. In contrast, female syncytotrophoblasts demonstrate upregulated expression of chemokine IL-10, this is thought to provide a safer invasion stage during blastocyst implantation. These differentially expressed genes (DEGs) in syncytotrophoblasts may be linked to downstream effects of genes that are associated with male or female CCs [1, 3 & 4]. Moreover, parental imprinting may contribute to these DEGs – where maternal imprinting of TNF-α and NF-κB in male placenta results in higher expression of paternal alleles, and paternal imprinting of IL-10 in female placenta leads to greater expression of the maternal alleles. The PAR1 and PAR2 genes in the petite arm of the Y chromosome demonstrated upregulation in male placenta, which is expected as only males would have the Yp PAR alleles [1, 3 & 4]. This coincides with the expression of Y-linked genes such as DDX3Y, EIF1AY and RPS4Y2 in chorionic villi. In this tissues they facilitate male-specific chromatin modifications and RNA splicing, which lead to the generation of H-Y antigens on MHC receptors of trophoblasts, stromal and Hofbauer cells [1, 3 & 4]. In comparison, X-linked genes that are upregulated in female placentas are XEs, where it gives a double of dose of 58 genes. These XE genes are thought to protect the female placenta against mitochondrial stressors such as poor maternal diets [1, 3 & 4].
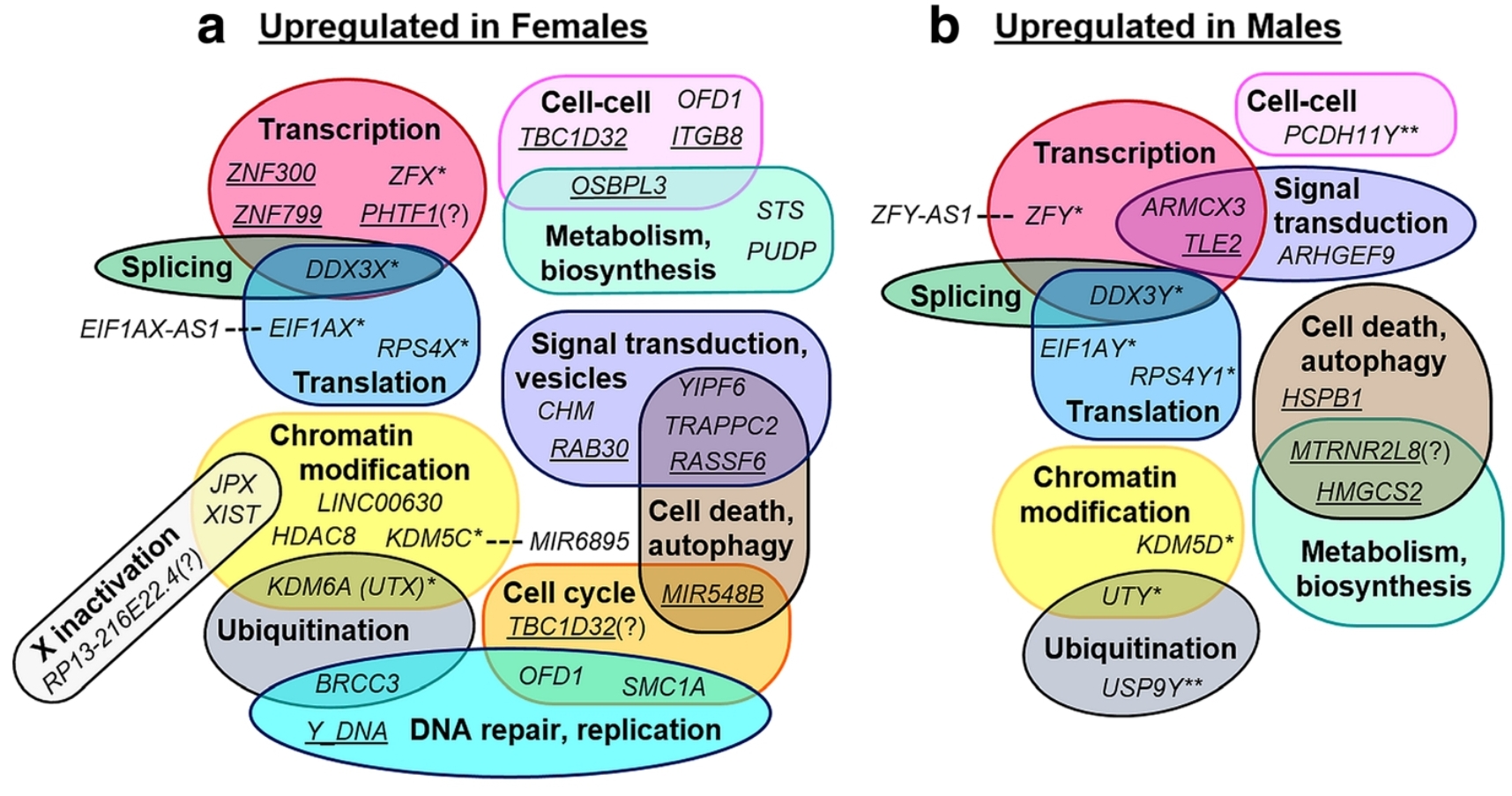
Figure 5 – Male vs Female Transcriptional Networks in Placenta. Most the genes in these separate transcriptional networks are not shared. Hence at a physiological level, male and female placentas behave different within similar contexts [1, 3 & 8]. In general, this is reflected in how male placentas implant faster and promote unfettered growth, whereas female placentas prioritize metabolic resilience against stressors and immune evasion. Additionally, this may explain why male placenta are more prone to recurrent miscarriages, pre-term birth, placental inflammation and spontaneous rupture of membranes; whereas, female placentas are more susceptible to foetal growth restriction, pre-term preeclampsia, small gestational size and placenta previa [1, 3 & 8]
The macro-anatomy and cell composition of the placenta are the very similar between males and females. However, male and female placentas have dinstinct physiological responses, in terms of hormone and chemokine signaling. Additionally, the individual cell types of the placenta may present with unknown micro-anotomical features that either male- or female-specific. These could be stained on placental samples with fluorescent antibody probes during immunohistochemistry. Comprehensive sequencing and quantitative PCR methods have been utilized to measure the sex differences in the dynamic placental transcriptomes. However, there has not been any established knock-down/gene editing studies that examined the effects of removing/reversing sex-biased gene expression profiles. These future studies could prove valuable in determining if all the observed physiological differences between male and female placenta are wholly caused by sex-specific transcriptomes. This venture may even provide more insight into the timing of these dynamic transcriptomes, with regards to performing the gene editing/knock-down animal/organoid models throughout multiple stages of gestation. Moreover, the sex-biased placental genes could be used in screening tests to detect adverse complications in gestating foetuses or pre-implanted embryos, and may be viable therapeutic targets for treating specific gestational complications.
References
- Braun, A. E., Mitchel, O. R., Gonzalez, T. L., Sun, T., Flowers, A. E., Pisarska, M. D., & Winn, V. D. (2022). Sex at the interface: The origin and impact of sex differences in the developing human placenta. Biology of Sex Differences, 13(1). https://doi.org/10.1186/s13293-022-00459-7
- Chuong, E. B. (2018). The placenta goes viral: Retroviruses control gene expression in pregnancy. PLOS Biology, 16(10). https://doi.org/10.1371/journal.pbio.3000028
- Gonzalez, T. L., Sun, T., Koeppel, A. F., Lee, B., Wang, E. T., Farber, C. R., Rich, S. S., Sundheimer, L. W., Buttle, R. A., Chen, Y.-D. I., Rotter, J. I., Turner, S. D., Williams, J., Goodarzi, M. O., & Pisarska, M. D. (2018). Sex differences in the late first trimester human placenta transcriptome. Biology of Sex Differences, 9(1). https://doi.org/10.1186/s13293-018-0165-y
- Inkster, A. M., Yuan, V., Konwar, C., Matthews, A. M., Brown, C. J., & Robinson, W. P. (2021). A cross-cohort analysis of autosomal DNA methylation sex differences in the term placenta. Biology of Sex Differences, 12(38). https://doi.org/10.1101/2021.03.08.434471
- Jansen, C. H., Kastelein, A. W., Kleinrouweler, C. E., Van Leeuwen, E., De Jong, K. H., Pajkrt, E., & Van Noorden, C. J. (2020). Development of placental abnormalities in location and anatomy. Acta Obstetricia et Gynecologica Scandinavica, 99(8), 983–993. https://doi.org/10.1111/aogs.13834
- Massimiani, M., Lacconi, V., La Civita, F., Ticconi, C., Rago, R., & Campagnolo, L. (2019). Molecular signaling regulating endometrium–blastocyst crosstalk. International Journal of Molecular Sciences, 21(1), 23. https://doi.org/10.3390/ijms21010023
- Ross, C., & Boroviak, T. E. (2020). Origin and function of the yolk sac in primate embryogenesis. Nature Communications, 11(3760). https://doi.org/10.1038/s41467-020-17575-w
- Soares, M. J., Varberg, K. M., & Iqbal, K. (2018). Hemochorial placentation: Development, function, and adaptations†. Biology of Reproduction, 99(1), 196–211. https://doi.org/10.1093/biolre/ioy049
- Sun, M., Wolf, G., Wang, Y., Ralls, S., Jin, J., Dunn-Fletcher, C. E., Muglia, L. J., & Macfarlan, T. S. (2020). Deciphering the evolution of the transcriptional and regulatory landscape in human placenta. bioRxiv. https://doi.org/10.1101/2020.09.11.289686


